Overview
Today we will learn to:
Use the six main data frame manipulation ‘verbs’ with pipes in dplyr
Understand how group_by() and summarize() can be combined to summarize datasets
Analyze a subset of data using logical filtering
Questions
How can I manipulate data frames without repeating myself?
The Problem with Base R
We often need to select observations, group data, or calculate statistics.
Using base R:
<- read.csv ("https://raw.githubusercontent.com/swcarpentry/r-novice-gapminder/main/episodes/data/gapminder_data.csv" mean (gapminder$ gdpPercap[gapminder$ continent == "Africa" ])mean (gapminder$ gdpPercap[gapminder$ continent == "Americas" ])mean (gapminder$ gdpPercap[gapminder$ continent == "Asia" ])
This is repetitive and error-prone!
The dplyr Package
The dplyr package provides functions for manipulating data frames that:
Reduce repetition
Reduce probability of errors
Are easier to read
Tidyverse
dplyr belongs to the Tidyverse - a family of R packages designed for data science.
These packages work harmoniously together. Learn more at: tidyverse.org
The Five Main Verbs
We’ll cover these commonly used functions:
select() - choose columnsfilter() - choose rowsgroup_by() - group datasummarize() - calculate summariesmutate() - create new columns
Plus the pipe operator: %>%
Loading dplyr
If not installed, run: install.packages("dplyr")
Using select()
Keep only specific columns:
<- select (gapminder, year, country, gdpPercap)head (year_country_gdp)
year country gdpPercap
1 1952 Afghanistan 779.4453
2 1957 Afghanistan 820.8530
3 1962 Afghanistan 853.1007
4 1967 Afghanistan 836.1971
5 1972 Afghanistan 739.9811
6 1977 Afghanistan 786.1134
Removing Columns with select()
Use - to remove columns:
<- select (gapminder, - continent)head (smaller_gapminder)
country year pop lifeExp gdpPercap
1 Afghanistan 1952 8425333 28.801 779.4453
2 Afghanistan 1957 9240934 30.332 820.8530
3 Afghanistan 1962 10267083 31.997 853.1007
4 Afghanistan 1967 11537966 34.020 836.1971
5 Afghanistan 1972 13079460 36.088 739.9811
6 Afghanistan 1977 14880372 38.438 786.1134
Introduction to Pipes
The pipe symbol %>% passes data from one function to the next:
<- gapminder %>% select (year, country, gdpPercap)head (year_country_gdp)
year country gdpPercap
1 1952 Afghanistan 779.4453
2 1957 Afghanistan 820.8530
3 1962 Afghanistan 853.1007
4 1967 Afghanistan 836.1971
5 1972 Afghanistan 739.9811
6 1977 Afghanistan 786.1134
Understanding Pipes
Step by step:
Start with gapminder data frame
Pass it using %>% to the next step
select() receives the data automatically
Fun Fact: Similar to the shell pipe | but the concept is the same!
Renaming Columns
Use rename() within a pipeline:
<- year_country_gdp %>% rename (gdp_per_capita = gdpPercap)head (tidy_gdp)
year country gdp_per_capita
1 1952 Afghanistan 779.4453
2 1957 Afghanistan 820.8530
3 1962 Afghanistan 853.1007
4 1967 Afghanistan 836.1971
5 1972 Afghanistan 739.9811
6 1977 Afghanistan 786.1134
Syntax: rename(new_name = old_name)
Using filter()
Keep only rows that meet criteria:
<- gapminder %>% filter (continent == "Europe" ) %>% select (year, country, gdpPercap)head (year_country_gdp_euro)
year country gdpPercap
1 1952 Albania 1601.056
2 1957 Albania 1942.284
3 1962 Albania 2312.889
4 1967 Albania 2760.197
5 1972 Albania 3313.422
6 1977 Albania 3533.004
Combining filter() Conditions
Filter with multiple conditions:
<- gapminder %>% filter (continent == "Europe" , year == 2007 ) %>% select (country, lifeExp)head (europe_lifeExp_2007)
country lifeExp
1 Albania 76.423
2 Austria 79.829
3 Belgium 79.441
4 Bosnia and Herzegovina 74.852
5 Bulgaria 73.005
6 Croatia 75.748
Challenge 1
Write a single command (which can span multiple lines and includes pipes) that will produce a data frame that has the African values for lifeExp, country and year, but not for other Continents.
How many rows does your data frame have and why?
Challenge 1 Solution
<- gapminder %>% filter (continent == "Africa" ) %>% select (year, country, lifeExp)nrow (year_country_lifeExp_Africa)
624 rows (52 African countries × 12 years of data)
Order of Operations
The order of operations matters!
If we used select() first, filter() would not be able to find the variable continent since we would have removed it.
Always filter before selecting when you need to filter on a column you won’t keep!
Using group_by()
Instead of filtering one group at a time, use group_by() to work with all groups:
'data.frame': 1704 obs. of 6 variables:
$ country : chr "Afghanistan" "Afghanistan" "Afghanistan" "Afghanistan" ...
$ year : int 1952 1957 1962 1967 1972 1977 1982 1987 1992 1997 ...
$ pop : num 8425333 9240934 10267083 11537966 13079460 ...
$ continent: chr "Asia" "Asia" "Asia" "Asia" ...
$ lifeExp : num 28.8 30.3 32 34 36.1 ...
$ gdpPercap: num 779 821 853 836 740 ...
group_by() Structure
str (gapminder %>% group_by (continent))
gropd_df [1,704 × 6] (S3: grouped_df/tbl_df/tbl/data.frame)
$ country : chr [1:1704] "Afghanistan" "Afghanistan" "Afghanistan" "Afghanistan" ...
$ year : int [1:1704] 1952 1957 1962 1967 1972 1977 1982 1987 1992 1997 ...
$ pop : num [1:1704] 8425333 9240934 10267083 11537966 13079460 ...
$ continent: chr [1:1704] "Asia" "Asia" "Asia" "Asia" ...
$ lifeExp : num [1:1704] 28.8 30.3 32 34 36.1 ...
$ gdpPercap: num [1:1704] 779 821 853 836 740 ...
- attr(*, "groups")= tibble [5 × 2] (S3: tbl_df/tbl/data.frame)
..$ continent: chr [1:5] "Africa" "Americas" "Asia" "Europe" ...
..$ .rows : list<int> [1:5]
.. ..$ : int [1:624] 25 26 27 28 29 30 31 32 33 34 ...
.. ..$ : int [1:300] 49 50 51 52 53 54 55 56 57 58 ...
.. ..$ : int [1:396] 1 2 3 4 5 6 7 8 9 10 ...
.. ..$ : int [1:360] 13 14 15 16 17 18 19 20 21 22 ...
.. ..$ : int [1:24] 61 62 63 64 65 66 67 68 69 70 ...
.. ..@ ptype: int(0)
..- attr(*, ".drop")= logi TRUE
Notice the grouped_df class - it’s now organized by continent!
Using summarize()
Combine group_by() with summarize() to calculate statistics per group:
<- gapminder %>% group_by (continent) %>% summarize (mean_gdpPercap = mean (gdpPercap))
# A tibble: 5 × 2
continent mean_gdpPercap
<chr> <dbl>
1 Africa 2194.
2 Americas 7136.
3 Asia 7902.
4 Europe 14469.
5 Oceania 18622.
Challenge 2
Calculate the average life expectancy per country.
Which has the longest average life expectancy and which has the shortest average life expectancy?
Challenge 2 Solution
<- gapminder %>% group_by (country) %>% summarize (mean_lifeExp = mean (lifeExp))%>% filter (mean_lifeExp == min (mean_lifeExp) | mean_lifeExp == max (mean_lifeExp))
# A tibble: 2 × 2
country mean_lifeExp
<chr> <dbl>
1 Iceland 76.5
2 Sierra Leone 36.8
Challenge 2 Solution (using arrange)
Another way using arrange():
# Shortest life expectancy %>% arrange (mean_lifeExp) %>% head (1 )
# A tibble: 1 × 2
country mean_lifeExp
<chr> <dbl>
1 Sierra Leone 36.8
# Longest life expectancy %>% arrange (desc (mean_lifeExp)) %>% head (1 )
# A tibble: 1 × 2
country mean_lifeExp
<chr> <dbl>
1 Iceland 76.5
Grouping by Multiple Variables
Group by more than one variable:
<- gapminder %>% group_by (continent, year) %>% summarize (mean_gdpPercap = mean (gdpPercap))head (gdp_bycontinents_byyear)
# A tibble: 6 × 3
# Groups: continent [1]
continent year mean_gdpPercap
<chr> <int> <dbl>
1 Africa 1952 1253.
2 Africa 1957 1385.
3 Africa 1962 1598.
4 Africa 1967 2050.
5 Africa 1972 2340.
6 Africa 1977 2586.
Multiple Summary Statistics
Create multiple summary variables at once:
<- gapminder %>% group_by (continent, year) %>% summarize (mean_gdpPercap = mean (gdpPercap),sd_gdpPercap = sd (gdpPercap),mean_pop = mean (pop),sd_pop = sd (pop)head (gdp_pop_bycontinents_byyear)
# A tibble: 6 × 6
# Groups: continent [1]
continent year mean_gdpPercap sd_gdpPercap mean_pop sd_pop
<chr> <int> <dbl> <dbl> <dbl> <dbl>
1 Africa 1952 1253. 983. 4570010. 6317450.
2 Africa 1957 1385. 1135. 5093033. 7076042.
3 Africa 1962 1598. 1462. 5702247. 7957545.
4 Africa 1967 2050. 2848. 6447875. 8985505.
5 Africa 1972 2340. 3287. 7305376. 10130833.
6 Africa 1977 2586. 4142. 8328097. 11585184.
count() and n()
Count observations per group with count():
%>% filter (year == 2002 ) %>% count (continent, sort = TRUE )
continent n
1 Africa 52
2 Asia 33
3 Europe 30
4 Americas 25
5 Oceania 2
Using n() in Calculations
Use n() for the number of observations in calculations:
%>% group_by (continent) %>% summarize (se_le = sd (lifeExp) / sqrt (n ()))
# A tibble: 5 × 2
continent se_le
<chr> <dbl>
1 Africa 0.366
2 Americas 0.540
3 Asia 0.596
4 Europe 0.286
5 Oceania 0.775
Multiple Summary Statistics Example
Calculate min, max, mean, and standard error:
%>% group_by (continent) %>% summarize (mean_le = mean (lifeExp),min_le = min (lifeExp),max_le = max (lifeExp),se_le = sd (lifeExp) / sqrt (n ())
# A tibble: 5 × 5
continent mean_le min_le max_le se_le
<chr> <dbl> <dbl> <dbl> <dbl>
1 Africa 48.9 23.6 76.4 0.366
2 Americas 64.7 37.6 80.7 0.540
3 Asia 60.1 28.8 82.6 0.596
4 Europe 71.9 43.6 81.8 0.286
5 Oceania 74.3 69.1 81.2 0.775
Using mutate()
Create new columns with mutate():
<- gapminder %>% mutate (gdp_billion = gdpPercap * pop / 10 ^ 9 ) %>% group_by (continent, year) %>% summarize (mean_gdpPercap = mean (gdpPercap),mean_gdp_billion = mean (gdp_billion)head (gdp_pop_bycontinents_byyear)
# A tibble: 6 × 4
# Groups: continent [1]
continent year mean_gdpPercap mean_gdp_billion
<chr> <int> <dbl> <dbl>
1 Africa 1952 1253. 5.99
2 Africa 1957 1385. 7.36
3 Africa 1962 1598. 8.78
4 Africa 1967 2050. 11.4
5 Africa 1972 2340. 15.1
6 Africa 1977 2586. 18.7
mutate() with ifelse()
Combine mutate() with ifelse() for conditional values:
<- gapminder %>% mutate (gdp_billion = ifelse (lifeExp > 25 , gdpPercap * pop / 10 ^ 9 , NA )) %>% group_by (continent, year) %>% summarize (mean_gdp_billion = mean (gdp_billion))head (gdp_pop_above25)
# A tibble: 6 × 3
# Groups: continent [1]
continent year mean_gdp_billion
<chr> <int> <dbl>
1 Africa 1952 5.99
2 Africa 1957 7.36
3 Africa 1962 8.78
4 Africa 1967 11.4
5 Africa 1972 15.1
6 Africa 1977 18.7
Scaling Values Conditionally
Update values based on conditions:
<- gapminder %>% mutate (gdp_futureExpectation = ifelse (lifeExp > 40 , gdpPercap * 1.5 , gdpPercap)) %>% group_by (continent, year) %>% summarize (mean_gdpPercap = mean (gdpPercap),mean_gdpPercap_expected = mean (gdp_futureExpectation)head (gdp_future)
# A tibble: 6 × 4
# Groups: continent [1]
continent year mean_gdpPercap mean_gdpPercap_expected
<chr> <int> <dbl> <dbl>
1 Africa 1952 1253. 1578.
2 Africa 1957 1385. 1803.
3 Africa 1962 1598. 2227.
4 Africa 1967 2050. 2953.
5 Africa 1972 2340. 3416.
6 Africa 1977 2586. 3830.
Combining dplyr and ggplot2
Load ggplot2:
Warning: package 'ggplot2' was built under R version 4.5.2
Traditional Approach
Creating an intermediate variable:
<- gapminder[gapminder$ continent == "Americas" , ]ggplot (data = americas, mapping = aes (x = year, y = lifeExp)) + geom_line () + facet_wrap (~ country) + theme (axis.text.x = element_text (angle = 45 ))
Piping into ggplot2
No intermediate variable needed:
%>% filter (continent == "Americas" ) %>% ggplot (mapping = aes (x = year, y = lifeExp)) + geom_line () + facet_wrap (~ country) + theme (axis.text.x = element_text (angle = 45 ))
Combining mutate and ggplot2
Create new columns and plot in one pipeline:
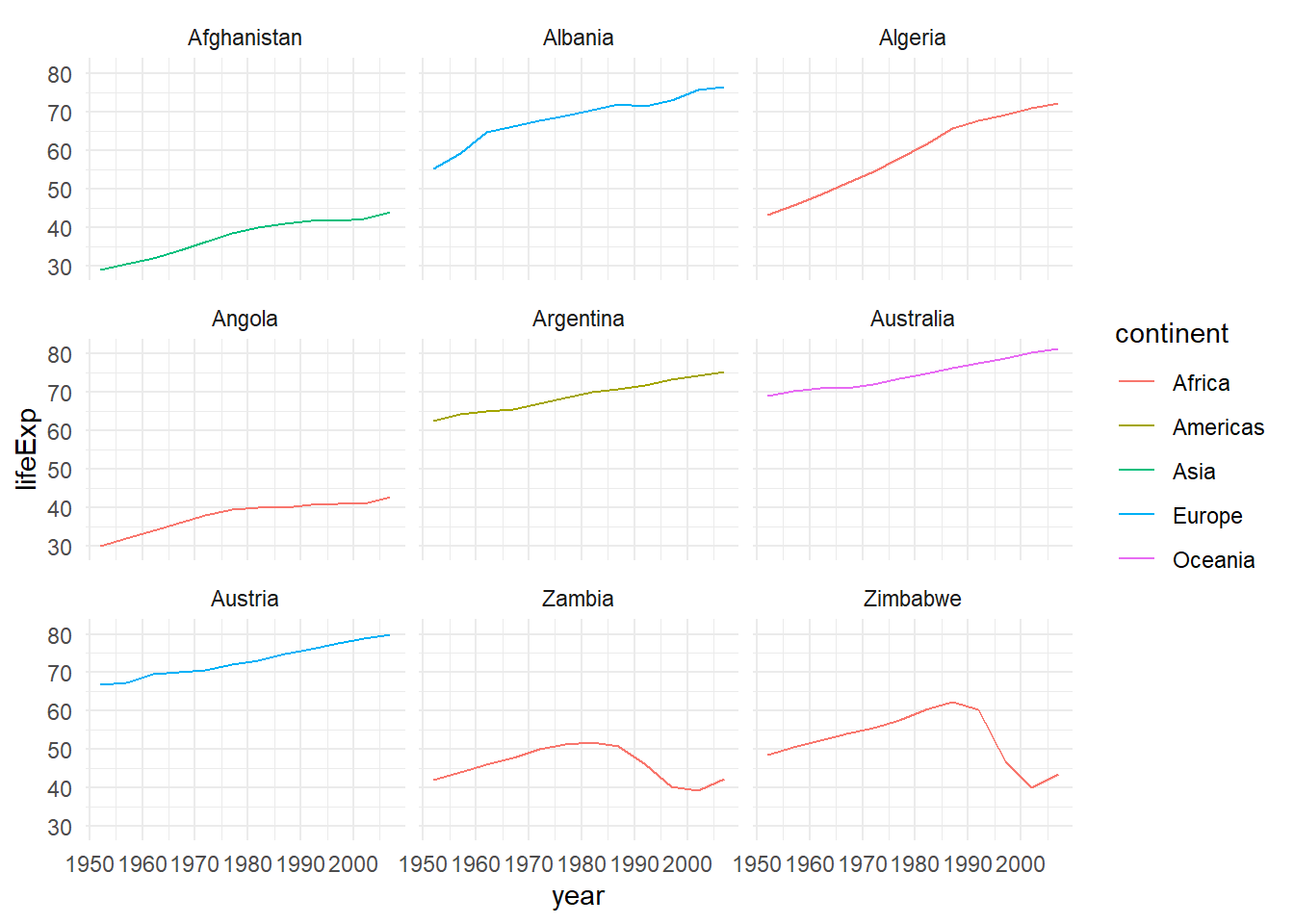
%>% mutate (startsWith = substr (country, 1 , 1 )) %>% filter (startsWith %in% c ("A" , "Z" )) %>% ggplot (aes (x = year, y = lifeExp, colour = continent)) + geom_line () + facet_wrap (vars (country)) + theme_minimal ()
Advanced Challenge
Calculate the average life expectancy in 2002 of 2 randomly selected countries for each continent.
Then arrange the continent names in reverse order.
Hint: Use the dplyr functions arrange() and sample_n().
Advanced Challenge Solution
set.seed (42 ) # For reproducibility <- gapminder %>% filter (year == 2002 ) %>% group_by (continent) %>% sample_n (2 ) %>% summarize (mean_lifeExp = mean (lifeExp)) %>% arrange (desc (mean_lifeExp))
# A tibble: 5 × 2
continent mean_lifeExp
<chr> <dbl>
1 Oceania 79.7
2 Asia 75.9
3 Europe 75.9
4 Americas 74.3
5 Africa 59.8
Key dplyr Verbs Summary
select()Choose columns
filter()Choose rows
group_by()Group data
summarize()Calculate summaries
mutate()Create new columns
arrange()Sort rows
count()Count observations
rename()Rename columns
Key Points
Use the dplyr package to manipulate data frames
Use select() to choose variables from a data frame
Use filter() to choose data based on values
Use group_by() and summarize() to work with subsets of data
Use mutate() to create new variables
Use pipes (%>%) to chain operations together
Important Reminders
Order of operations matters when combining select() and filter()
group_by() changes the data frame structuren() returns the count of observations in the current groupPipes make code more readable and reduce intermediate variables
Combine dplyr with ggplot2 for powerful data visualization workflows